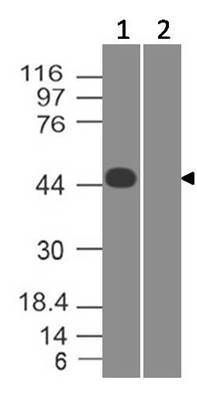
Coronavirus (COVID-19) Spike Antibody
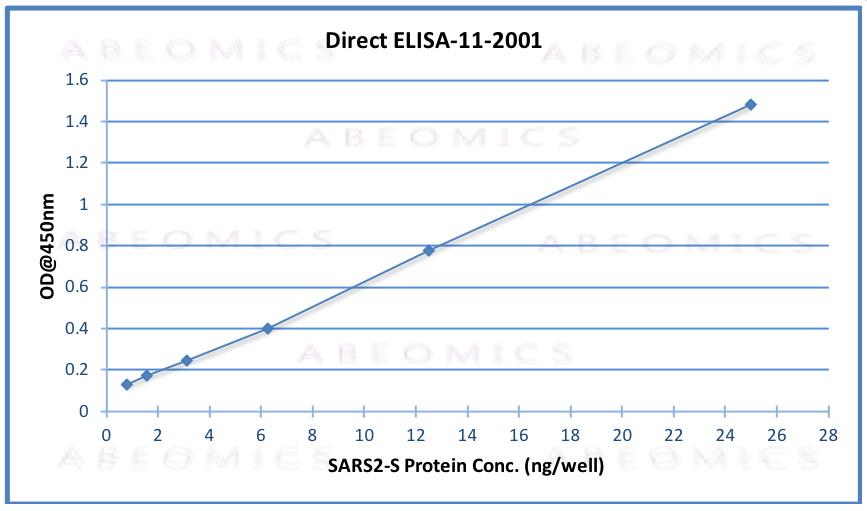
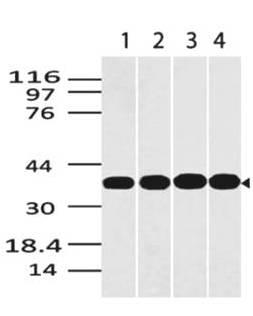
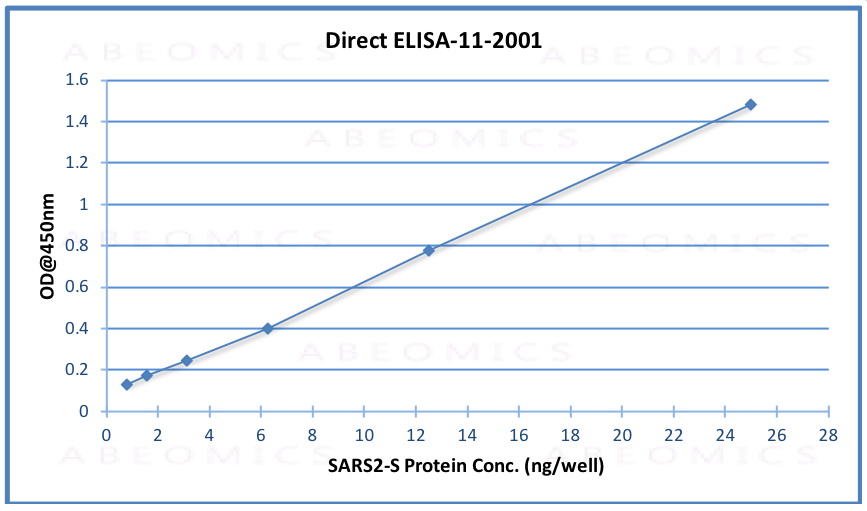
Fig.1: Wells of a 96-microtiter plate were coated with different concentration of a mammalian expressed full-length SARS-Cov-2/nCov/COVID-19 Spike protein. The binding was detected by addition of 200 ng 11-2001 polyclonal per well. The reactivity was detected by a HRP-conjugated goat-anti-rabbit polyclonal antibody.
Roll over image to zoom in
Shipping Info:
Order now and get it on Friday June 05, 2026
Same day delivery FREE on San Diego area orders placed by 1.00 PM
| Format : | Purified |
| Amount : | 100 µg |
| Isotype : | Rabbit IgG |
| Purification : | Protein A Purification |
| Content : | 25 µg in 50 µl/100 µg in 200 µl PBS containing 0.05% BSA and 0.05% sodium azide. Sodium azide is highly toxic. |
| Storage condition : | Store the antibody at 4°C, stable for 6 months. For long-term storage, store at -20°C. Avoid repeated freeze and thaw cycles. |
| Uniprot ID : | P0DTC2 |
| Alternative Name : | nCov, Sars-Cov-2 |
| Immunogen Information : | A partial length recombinant coronavirus Spike Protein (amino acids 970-985) was used as the immunogen for this antibody. |
The spike (S) protein of nCoV/SARS-CoV-2/COVID-19 is one of the structural glycoproteins that remains embedded in viral envelope and acts as the fundamental component of early viral infection of nCoV/SARS-CoV-2/COVID-19 upon binding the host receptor. The nCoV/SARS-CoV-2/COVID-19 has a trimeric spike protein which has two main domains such as S1 domain for receptor binding and S2 domain for membrane fusion and several specific cleavage sites in S1- S2 boundary junction that needs a novel, endocytic protease- primed cleavage to get activated during infection. It mainly binds to the furin protein on the cell membrane which performs trypsin like proteolytic cleavage and then the protein gets activated facilitating its entry into the host. This transmembrane spike protein of nCoV/SARS-CoV-2/COVID-19 shares binding property to the Angiotensin Converting Enzyme 2 (ACE2) likely to that of SARS- CoV. The high affinity of nCoV/SARS-CoV-2/COVID-19 Spike protein for human ACE2 may contribute to the apparent ease with which nCoV/SARS-CoV-2/COVID-19 can spread from human-to-human and make nCoV/SARS-CoV-2/COVID-19 pandemic.
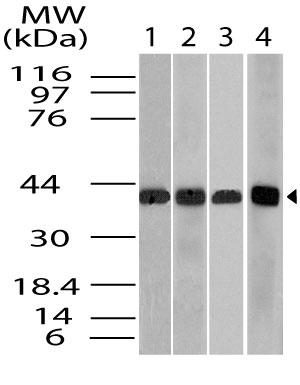
Recommended dilutions: WB: 2-4 µg/ml. However, this need to be optimized based on the research applications.
For Research Use Only. Not for use in diagnostic/therapeutics procedures.
1. Minimal system for assembly of SARS-CoV-2 virus like particles: Heather Swann, Abhimanyu Sharma, Benjamin Preece, Abby Peterson, Crystal Eldridge, David M. Belnap, Michael Vershinin, Saveez Saffarian bioRxiv 2020.06.01.128058; doi: https://doi.org/10.1101/2020.06.01.128058
|
There are currently no product reviews
|




.png)