EGFL6 Mouse
Shipping Info:
For estimated delivery dates, please contact us at [email protected]
| Amount : | 10 µg |
| Purification : | Greater than 95.0% as determined by SDS-PAGE. |
| Content : | EGFL6 protein solution ( 0.5mg/ml ) contains Phosphate Buffered Saline (pH 7.4) containing 10% glycerol. |
| Storage condition : | Store at 4°C if entire vial will be used within 2-4 weeks. Store, frozen at -20°C for longer periods of time. For long term storage it is recommended to add a carrier protein (0.1% HSA or BSA).Avoid multiple freeze-thaw cycles. |
| AA sequence : | ADLTMKKKVK LKMVTPRPAS TRVPKVNLPY SSEEGVSRGR NYDGEQKKKE EGKRERLEEEKGEKTLRNEV EQERTLRGDV FSPKVNEAED LDLVYVQRKE LNSKLKHKDL NISVDCSFDLGVCDWKQDRE DDFDWHPADR DNDVGYYMAV PALAGHKKNI GRLKLLLPNL TPQSNFCLLFDYRLAGDKVG KLRVFVKNSN NALAWEETKN EDGRWRTGKI QLYQGIDTTK SVIFEAERGK GKTGEIAVDG VLLVSGLCPD DFLSVEGHHH HHH. |
| Alternative Name : | Epidermal growth factor-like protein 6, EGF-L6, Egfl6, Maeg. |
Source: Sf9, Insect cells.
Sterile filtered colorless solution.
Epidermal Growth FactorÂlike Domain Multiple 6 (EGFL6) belongs to the EGF repeat superfamily of proteins, whose members are involved in the regulation of cell cycle, proliferation, and developmental processes. EGFL6 gene product contains a signal peptide, suggesting that EGFL6 is secreted; an EGF repeat region consisting of four complete EGF-like repeats and 1 partial EGF-like repeat, 3 of which have a calcium-binding consensus sequence; an arg-gly-asp integrin association motif; and a MAM domain, which is assumed to have an adhesive function. Within shared regions, human EGFL6 shares 75% and 78% amino acid sequence identity with the mouse and rat orthologs, respectively. EGFL6 is expressed in various fetal tissues during early development such as the lung, heart, liver, spleen, cochlea and the placenta, as well as meningioma tumors.
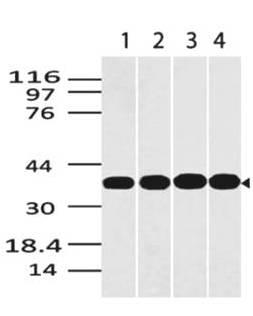
EGFL6 Mouse Recombinant produced in Sf9 Insect cells is a single, glycosylated polypeptide chain containing 273 amino acids (287-550a.a.) and having a molecular mass of 31.1kDa (Molecular size on SDS-PAGE will appear at approximately 28-40kDa).EGFL6 is expressed with a 9 amino acid His tag at C-Terminus and purified by proprietary chromatographic techniques.
Sterile filtered colorless solution.
Epidermal Growth FactorÂlike Domain Multiple 6 (EGFL6) belongs to the EGF repeat superfamily of proteins, whose members are involved in the regulation of cell cycle, proliferation, and developmental processes. EGFL6 gene product contains a signal peptide, suggesting that EGFL6 is secreted; an EGF repeat region consisting of four complete EGF-like repeats and 1 partial EGF-like repeat, 3 of which have a calcium-binding consensus sequence; an arg-gly-asp integrin association motif; and a MAM domain, which is assumed to have an adhesive function. Within shared regions, human EGFL6 shares 75% and 78% amino acid sequence identity with the mouse and rat orthologs, respectively. EGFL6 is expressed in various fetal tissues during early development such as the lung, heart, liver, spleen, cochlea and the placenta, as well as meningioma tumors.
EGFL6 Mouse Recombinant produced in Sf9 Insect cells is a single, glycosylated polypeptide chain containing 273 amino acids (287-550a.a.) and having a molecular mass of 31.1kDa (Molecular size on SDS-PAGE will appear at approximately 28-40kDa).EGFL6 is expressed with a 9 amino acid His tag at C-Terminus and purified by proprietary chromatographic techniques.
|
There are currently no product reviews
|








.png)