VCAM1 Human, sf9
Shipping Info:
For estimated delivery dates, please contact us at [email protected]
| Amount : | 10 µg |
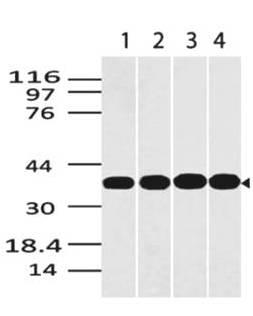
| Purification : | Greater than 90.0% as determined by SDS-PAGE. |
| Content : | VCAM1 protein solution (0.5mg/ml) contains Phosphate Buffered Saline (pH 7.4) and 10% glycerol. |
| Storage condition : | Store at 4°C if entire vial will be used within 2-4 weeks. Store, frozen at -20°C for longer periods of time. For long term storage it is recommended to add a carrier protein (0.1% HSA or BSA). Avoid multiple freeze-thaw cycles. |
| AA sequence : | ADPEFFKIET TPESRYLAQI GDSVSLTCST TGCESPFFSW RTQIDSPLNG KVTNEGTTST LTMNPVSFGN EHSYLCTATC ESRKLEKGIQ VEIYSFPKDP EIHLSGPLEA GKPITVKCSV ADVYPFDRLE IDLLKGDHLM KSQEFLEDAD RKSLETKSLE VTFTPVIEDI GKVLVCRAKL HIDEMDSVPT VRQAVKELQV YISPKNTVIS VNPSTKLQEG GSVTMTCSSE GLPAPEIFWS KKLDNGNLQH LSGNATLTLI AMRMEDSGIY VCEGVNLIGK NRKEVELIVQ EKPFTVEISP GPRIAAQIGD SVMLTCSVMG CESPSFSWRT QIDSPLSGKV RSEGTNSTLT LSPVSFENEH SYLCTVTCGH KKLEKGIQVE LYSFPRDPEI EMSGGLVNGS SVTVSCKVPS VYPLDRLEIE LLKGETILEN IEFLEDTDMK SLENKSLEMT FIPTIEDTGK ALVCQAKLHI DDMEFEPKQR QSTQTLYVNV APRDTTVLVS PSSILEEGSS VNMTCLSQGF PAPKILWSRQ LPNGELQPLS ENATLTLIST KMEDSGVYLC EGINQAGRSR KEVELIIQVT PKDIKLTAFP SESVKEGDTV IISCTCGNVP ETWIILKKKA ETGDTVLKSI DGAYTIRKAQ LKDAGVYECE SKNKVGSQLR SLTLDVQGRE NNKDYFSPEH HHHHH. |
| Alternative Name : | Vascular Cell Adhesion Molecule 1, CD106 Antigen, INCAM-100 , Vascular Cell Adhesion Protein 1, V-CAM 1, VCAM-1, CD106, L1CAM. |
Source: Sf9, Baculovirus cells.
Sterile Filtered colorless solution.
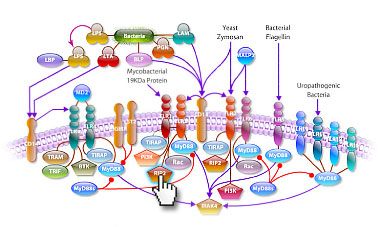
VCAM1 is a member of the Ig superfamily. It is a cell surface sialoglycoprotein expressed by cytokine activated endothelium. VCAM-1 contains 6 or 7 immunoglobulin domains, and is expressed on both large and small vessels only after the endothelial cells are stimulated by cytokines. The protein has a number of functions including the regulation of leukocyte migration, leukocyte endothelial cell adhesion and signal transduction and may play a role in a number of inflammatory diseases (artherosclerosis and rheumatoid arthritis).
VCAM1 produced in Sf9 Baculovirus cells is a single, glycosylated polypeptide chain containing 685 amino acids (25-698 a.a.) and having a molecular mass of 75.6kDa (Molecular size on SDS-PAGE will appear at approximately 70-100kDa). VCAM1 is expressed with a 11 amino acid His tag at C-Terminus and purified by proprietary chromatographic techniques.
Sterile Filtered colorless solution.
VCAM1 is a member of the Ig superfamily. It is a cell surface sialoglycoprotein expressed by cytokine activated endothelium. VCAM-1 contains 6 or 7 immunoglobulin domains, and is expressed on both large and small vessels only after the endothelial cells are stimulated by cytokines. The protein has a number of functions including the regulation of leukocyte migration, leukocyte endothelial cell adhesion and signal transduction and may play a role in a number of inflammatory diseases (artherosclerosis and rheumatoid arthritis).
VCAM1 produced in Sf9 Baculovirus cells is a single, glycosylated polypeptide chain containing 685 amino acids (25-698 a.a.) and having a molecular mass of 75.6kDa (Molecular size on SDS-PAGE will appear at approximately 70-100kDa). VCAM1 is expressed with a 11 amino acid His tag at C-Terminus and purified by proprietary chromatographic techniques.
|
There are currently no product reviews
|




![Anti-p53 (Phospho-Ser392) Monoclonal Antibody (Clone:FP3.2 [FPS392]) Anti-p53 (Phospho-Ser392) Monoclonal Antibody (Clone:FP3.2 [FPS392])](https://media.abeomics.com/images/30-1325/1.jpg)

.png)