ST6GAL1 Human
Shipping Info:
For estimated delivery dates, please contact us at [email protected]
| Amount : | 20 µg |
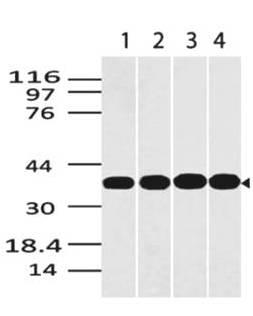
| Purification : | Greater than 90.0% as determined by SDS-PAGE. |
| Content : | ST6GAL1 protein solution (1mg/ml) containing 20mM Tris-HCl (pH 8.0) and 10% glycerol. |
| Storage condition : | Store at 4°C if entire vial will be used within 2-4 weeks. Store, frozen at -20°C for longer periods of time. For long term storage it is recommended to add a carrier protein (0.1% HSA or BSA).Avoid multiple freeze-thaw cycles. |
| AA sequence : | MGSSHHHHHH SSGLVPRGSH MGSKEKKKGS YYDSFKLQTK EFQVLKSLGK LAMGSDSQSV SSSSTQDPHR GRQTLGSLRG LAKAKPEASF QVWNKDSSSK NLIPRLQKIW KNYLSMNKYK VSYKGPGPGI KFSAEALRCH LRDHVNVSMV EVTDFPFNTS EWEGYLPKES IRTKAGPWGR CAVVSSAGSL KSSQLGREID DHDAVLRFNG APTANFQQDV GTKTTIRLMN SQLVTTEKRF LKDSLYNEGI LIVWDPSVYH SDIPKWYQNP DYNFFNNYKT YRKLHPNQPF YILKPQMPWE LWDILQEISP EEIQPNPPSS GMLGIIIMMT LCDQVDIYEF LPSKRKTDVC YYYQKFFDSA CTMGAYHPLL YEKNLVKHLN QGTDEDIYLL GKATLPGFRT IHC |
| Alternative Name : | ST6 Beta-Galactosamide Alpha-2,6-Sialyltranferase 1, ST6Gal I, SIAT1, CMP-N-Acetylneuraminate-Beta-Galactosamide-Alpha-2,6-Sialyltransferase 1, B-Cell Antigen CD75, Alpha 2,6-ST 1, EC 2.4.99.1, ST6GalI, CMP-N-Acetylneuraminate Beta-Galactosamide Alpha-2,6-Sialyltransferase, Sialyltransferase 1 (Beta-Galactoside Alpha-2,6-Sialyltransferase), Sialyltransferase 1 (Beta-Galactoside Alpha-2,6-Sialytransferase),Beta-Galactoside Alpha-2,6-Sialyltransferase 1, Sialyltransferase 1, ST6N, ST6GAL1. |
Source: Escherichia Coli.
Sterile Filtered colorless solution.
ST6GAL1 also known as ST6 Beta-Galactosamide Alpha-2,6-Sialyltranferase 1, is part of the glycosyltransferase family 29. ST6GAL1 is a type II membrane protein which catalyzes the transfer of sialic acid from CMP-sialic acid to galactose-containing substrates. Furthermore, ST6GAL1 is normally found in the Golgi however it can be proteolytically processed to a soluble form, ST6GAL1 is also involved in the generation of the cell-surface carbohydrate determinants as well differentiation antigens HB-6, CD75, and CD76.Â
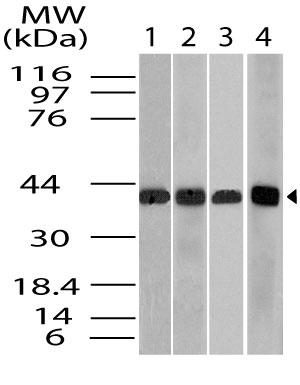
ST6GAL1 Human Recombinant produced in E.Coli is a single, non-glycosylated polypeptide chain containing 403 amino acids (27-406 a.a) and having a molecular mass of 46kDa.ST6GAL1 is fused to a 23 amino acid His-tag at N-terminus & purified by proprietary chromatographic techniques.
Sterile Filtered colorless solution.
ST6GAL1 also known as ST6 Beta-Galactosamide Alpha-2,6-Sialyltranferase 1, is part of the glycosyltransferase family 29. ST6GAL1 is a type II membrane protein which catalyzes the transfer of sialic acid from CMP-sialic acid to galactose-containing substrates. Furthermore, ST6GAL1 is normally found in the Golgi however it can be proteolytically processed to a soluble form, ST6GAL1 is also involved in the generation of the cell-surface carbohydrate determinants as well differentiation antigens HB-6, CD75, and CD76.Â
ST6GAL1 Human Recombinant produced in E.Coli is a single, non-glycosylated polypeptide chain containing 403 amino acids (27-406 a.a) and having a molecular mass of 46kDa.ST6GAL1 is fused to a 23 amino acid His-tag at N-terminus & purified by proprietary chromatographic techniques.
|
There are currently no product reviews
|





.png)